simdkalman documentation¶
Fast Kalman filters in Python leveraging single-instruction multiple-data vectorization. That is, running n similar Kalman filters on n independent series of observations. Not to be confused with SIMD processor instructions.
See full documentation.
Installation: pip install simdkalman
Source code: https://github.com/oseiskar/simdkalman
License: MIT
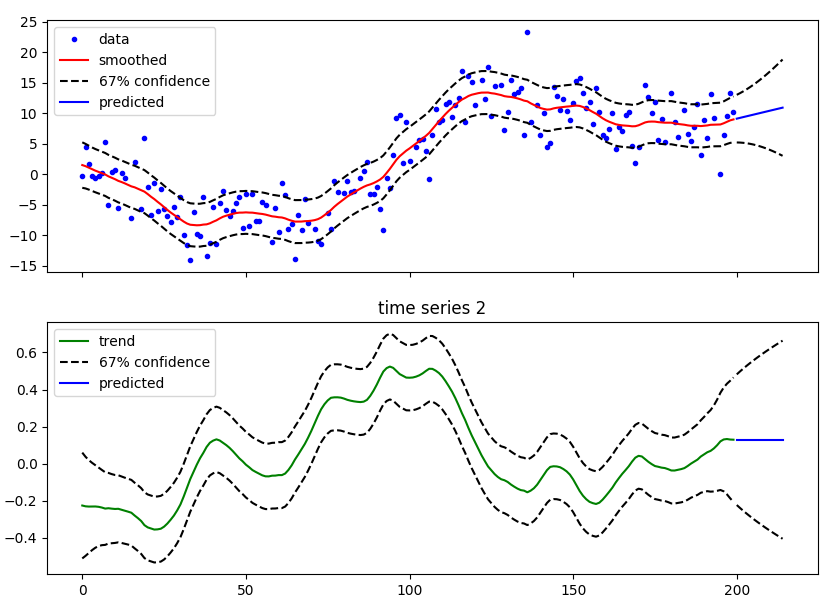
Terminology¶
Following the notation in [1], the Kalman filter framework consists of a dynamic model (state transition model)
and a measurement model (observation model)
where the vector \(x\) is the (hidden) state of the system and \(y\) is an observation. A and H are matrices of suitable shape and \(Q\), \(R\) are positive-definite noise covariance matrices.
Usage example¶
Define model
import simdkalman
import numpy as np
kf = simdkalman.KalmanFilter(
state_transition = [[1,1],[0,1]], # matrix A
process_noise = np.diag([0.1, 0.01]), # Q
observation_model = np.array([[1,0]]), # H
observation_noise = 1.0) # R
Generate some fake data
import numpy.random as random
# 100 independent time series
data = random.normal(size=(100, 200))
# with 10% of NaNs denoting missing values
data[random.uniform(size=data.shape) < 0.1] = np.nan
Smooth all data
smoothed = kf.smooth(data,
initial_value = [1,0],
initial_covariance = np.eye(2) * 0.5)
# second timeseries, third time step, hidden state x
print('mean')
print(smoothed.states.mean[1,2,:])
print('covariance')
print(smoothed.states.cov[1,2,:,:])
mean
[ 0.29311384 -0.06948961]
covariance
[[ 0.19959416 -0.00777587]
[-0.00777587 0.02528967]]
Predict new data for a single series (1d case)
predicted = kf.predict(data[1,:], 123)
# predicted observation y, third new time step
pred_mean = predicted.observations.mean[2]
pred_stdev = np.sqrt(predicted.observations.cov[2])
print('%g +- %g' % (pred_mean, pred_stdev))
1.71543 +- 1.65322
For complete code examples with figures, see: https://github.com/oseiskar/simdkalman/blob/master/examples/ and this Gist.
Usage of multi-dimensional observations is demonstrated in this example.
Extended Kalman Filters (EKF) can be implemented using the simdkalman.primitives module, see
example.
Class documentation¶
- class simdkalman.KalmanFilter(state_transition, process_noise, observation_model, observation_noise)¶
The main Kalman filter class providing convenient interfaces to vectorized smoothing and filtering operations on multiple independent time series.
As long as the shapes of the given parameters match reasonably according to the rules of matrix multiplication, this class is flexible in their exact nature accepting
scalars:
process_noise = 0.1(2d) numpy matrices:
process_noise = numpy.eye(2)2d arrays:
observation_model = [[1,2]]3d arrays and matrices for vectorized computations. Unlike the other options, this locks the shape of the inputs that can be processed by the smoothing and prediction methods.
- Parameters:
state_transition – State transition matrix \(A\)
process_noise – Process noise (state transition covariance) matrix \(Q\)
observation_model – Observation model (measurement model) matrix \(H\)
observation_noise – Observation noise (measurement noise covariance) matrix \(R\)
- compute(data, n_test, initial_value=None, initial_covariance=None, smoothed=True, filtered=False, states=True, covariances=True, observations=True, likelihoods=False, gains=False, log_likelihood=False, verbose=False)¶
Smoothing, filtering and prediction at the same time. Used internally by other methods, but can also be used directly if, e.g., both smoothed and predicted data is wanted.
See smooth and predict for explanation of the common parameters. With this method, there also exist the following flags.
- Parameters:
smoothed (boolean) – compute Kalman smoother (used by smooth)
filtered (boolean) – return (one-way) filtered data
likelihoods (boolean) – return likelihoods of each step
gains (boolean) – return Kalman gains and pairwise covariances (used by the EM algorithm). If true, the gains are provided as a member of the relevant subresult
filtered.gainsand/orsmoothed.gains.log_likelihood (boolean) – return the log-likelihood(s) for the entire series. If matrix data is given, this will be a vector where each element is the log-likelihood of a single row.
- Return type:
result object whose fields depend on of the above parameter flags are True. The possible values are:
smoothed(the return value of smooth, may containsmoothed.gains),filtered(likesmoothed, may also containfiltered.gains),predicted(the return value of predict ifn_test > 0)pairwise_covariances,likelihoodsandlog_likelihood.
- predict(data, n_test, initial_value=None, initial_covariance=None, states=True, observations=True, covariances=True, verbose=False)¶
Filter past data and predict a given number of future values. The data can be given as either of
1d array, like
[1,2,3,4]. In this case, one Kalman filter is used and the return value structure will contain an 1d array ofobservations(both.meanand.covwill be 1d).2d matrix, whose each row is interpreted as an independent time series, all of which are filtered independently. The returned
observationsmembers will be 2-dimensional in this case.3d matrix, whose the last dimension can be used for multi-dimensional observations, i.e,
data[1,2,:]defines the components of the third observation of the second series. In the-multi-dimensional case the returnedobservations.meanwill be 3-dimensional andobservations.cov4-dimensional.
Initial values and covariances can be given as scalars or 2d matrices in which case the same initial states will be used for all rows or 3d arrays for different initial values.
- Parameters:
data – Past data
n_test (integer) – number of future steps to predict.
initial_value – Initial value \({\mathbb E}[x_0]\)
initial_covariance – Initial uncertainty \({\rm Cov}[x_0]\)
states (boolean) – predict states \(x\)?
observations (boolean) – predict observations \(y\)?
covariances (boolean) – include covariances in predictions?
- Return type:
Result object with fields
statesandobservations, if the respective parameter flags are set to True. Both areGaussianresult objects with fieldsmeanandcov(if the covariances flag is True)
- predict_next(m, P)¶
Single prediction step
- Parameters:
m – \({\mathbb E}[x_{j-1}]\), the previous mean
P – \({\rm Cov}[x_{j-1}]\), the previous covariance
- Return type:
(prior_mean, prior_cov)predicted mean and covariance \({\mathbb E}[x_j]\), \({\rm Cov}[x_j]\)
- predict_observation(m, P)¶
Probability distribution of observation \(y\) for a given distribution of \(x\)
- Parameters:
m – \({\mathbb E}[x]\)
P – \({\rm Cov}[x]\)
- Return type:
mean \({\mathbb E}[y]\) and covariance \({\rm Cov}[y]\)
- smooth(data, initial_value=None, initial_covariance=None, observations=True, states=True, covariances=True, verbose=False)¶
Smooth given data, which can be either
1d array, like
[1,2,3,4]. In this case, one Kalman filter is used and the return value structure will contain an 1d array ofobservations(both.meanand.covwill be 1d).2d matrix, whose each row is interpreted as an independent time series, all of which are smoothed independently. The returned
observationsmembers will be 2-dimensional in this case.3d matrix, whose the last dimension can be used for multi-dimensional observations, i.e,
data[1,2,:]defines the components of the third observation of the second series. In the-multi-dimensional case the returnedobservations.meanwill be 3-dimensional andobservations.cov4-dimensional.
Initial values and covariances can be given as scalars or 2d matrices in which case the same initial states will be used for all rows or 3d arrays for different initial values.
- Parameters:
data – 1d or 2d data, see above
initial_value – Initial value \({\mathbb E}[x_0]\)
initial_covariance – Initial uncertainty \({\rm Cov}[x_0]\)
states (boolean) – return smoothed states \(x\)?
observations (boolean) – return smoothed observations \(y\)?
covariances (boolean) – include covariances results? For lower memory usage, set this flag to
False.
- Return type:
Result object with fields
statesandobservations, if the respective parameter flags are set to True. Both areGaussianresult objects with fieldsmeanandcov(if the covariances flag is True)
- smooth_current(m, P, ms, Ps)¶
Simgle Kalman smoother backwards step
- Parameters:
m – \({\mathbb E}[x_j|y_1,\ldots,y_j]\), the filtered mean of \(x_j\)
P – \({\rm Cov}[x_j|y_1,\ldots,y_j]\), the filtered covariance of \(x_j\)
ms – \({\mathbb E}[x_{j+1}|y_1,\ldots,y_T]\)
Ps – \({\rm Cov}[x_{j+1}|y_1,\ldots,y_T]\)
- Return type:
(smooth_mean, smooth_covariance, smoothing_gain)smoothed mean \({\mathbb E}[x_j|y_1,\ldots,y_T]\), and covariance \({\rm Cov}[x_j|y_1,\ldots,y_T]\) & smoothing gain \(C\)
- update(m, P, y, log_likelihood=False)¶
Single update step with NaN check.
- Parameters:
m – \({\mathbb E}[x_j|y_1,\ldots,y_{j-1}]\), the prior mean of \(x_j\)
P – \({\rm Cov}[x_j|y_1,\ldots,y_{j-1}]\), the prior covariance of \(x_j\)
y – observation \(y_j\)
log_likelihood – compute log-likelihood?
- Return type:
(posterior_mean, posterior_covariance, log_likelihood)posterior mean \({\mathbb E}[x_j|y_1,\ldots,y_j]\) & covariance \({\rm Cov}[x_j|y_1,\ldots,y_j]\) and, if requested, log-likelihood. If \(y_j\) is NaN, returns the prior mean and covariance instead
Primitives¶
The simdkalman.primitives module contains low-level Kalman filter computation
steps with multi-dimensional input arrays. See this page
for full documentation.